Introduction
Modern wearable muscle near-infrared spectroscopy (mNIRS) devices make it easier than ever to monitor local muscle oxygenation during dynamic activities.
The real challenge comes with deciding how to clean, filter, process, and eventually interpret those data.
The mnirs package aims to provide standardised, reproducible methods for reading, processing, and analysing mNIRS data. The goal is to help practitioners detect meaningful signal from noise, and improve our confidence in interpreting and applying information to the clients we work with.
In this vignette we will demonstrate how to:
📂 Read data files exported from commercial wearable NIRS devices, and import NIRS channels into a standard data frame format with metadata, ready for further processing.
📊 Plot and visualise data frames of class
"mnirs".🔍 Retrieve metadata stored with data frames of class
"mnirs"to avoid repetitively specifying which channels to process.⏱️ Resample data to a higher or lower sample rate, to correct irregular sampling periods, or to match the frequency of other data sources for synchronisation.
🧹 Detect and replace local outliers, invalid values, and interpolate across missing data.
📈️ Apply digital filtering to optimise signal-to-noise ratio for the responses observed in our data.
⚖️ Shift and rescale across multiple NIRS channels, to normalise signal dynamic range while preserving absolute or relative scaling between muscle sites and NIRS channels.
🔀 Demonstrate a complete data wrangling process using pipe-friendly functions.
🧮 Detect and extract intervals for further analysis.
mnirs is designed to process NIRS data, but it can be used to read, clean, and process other time series data which require many of the same processing steps. Enjoy!
📂 Read data from file
We will read an example data file with two NIRS channels from an incremental ramp cycling assessment recorded with Moxy muscle oxygen monitor.
First, install and load the mnirs package and other required libraries.
mnirs can be installed with install.packages("mnirs").
The first function called will often be read_mnirs(). This is used to read data from .csv or .xls(x) files exported from common wearable NIRS devices.
Exported data files will often have multiple rows of file header metadata before the data table with NIRS recordings begins. read_mnirs() can extract and return this data table along with the file metadata for further processing and analysis.
See read_mnirs() for more details.
read_mnirs()
-
file_pathSpecify the location of the data file, including file extension. e.g.
"./my_file.xlsx"or"C:/myfolder/my_file.csv".
Example data files
A few example data files are included in the
mnirspackage. File paths can be accessed withexample_mnirs().
-
nirs_channelsMultiple NIRS channels can be specified from the data file as a vector of names. If no
nirs_channelsare specified,read_mnirs()will attempt to recognise the NIRS device file format, and return the full data frame with all detected columns. This can be useful for file exploration, to find the target channel names. However, best practice is to specify the desirednirs_channelsexplicitly. -
time_channelA time or sample channel name from the data table can be specified. If left blank, the function will attempt to identify the time column automatically, however, best practice is to specify the
time_channelexplicitly. -
event_channelOptionally, a channel can be specified which indicates character event labels or integer lap values in the data table.
These channel names are used to detect the data table within the file, and must match exactly with text strings in the file on the same row. We can rename these channels when reading data by specifying a named character vector:
nirs_channels = c(
renamed1 = "original_name1",
renamed2 = "original_name2"
)-
sample_rateThe sample rate (in Hz) of the exported data can either be specified explicitly, or it will be estimated from
time_channel.
Automatic detection usually works well unless there is irregular sampling, or thetime_channelis a count of samples rather than a time value. For example, Oxysoft exports a column of sample numbers rather than time values. For Oxysoft files specifically, the function will recognise and read the correct sample rate from the file metadata. However, in most casessample_rateshould be defined explicitly if known. -
add_timestampFALSEby default; iftime_channelis in date-time (POSIXct) format (e.g., hh:mm:ss), by default it will be converted to numeric time in seconds.
Ifadd_timestamp = TRUE, the date-time start time value intime_channelor in the file metadata will be extracted and a"timestamp"column will be added to the returned data frame. This can be useful for synchronising devices based on system time. -
zero_timeFALSEby default; iftime_channelvalues start at a non-zero value,zero_time = TRUEwill re-calculate time starting from zero. Iftime_channelwas converted from date-time format, it will always be re-calculated from zero regardless of this option. -
keep_allFALSEby default; only the channels explicitly specified will be returned in a data frame.keep_all = TRUEwill return all columns detected from the data table in the file.
Blank/empty columns will be omitted. Duplicate column names will be repaired by appending a suffix"_n", and empty column names will be renamed ascol_n; wherenis equal to the column number in the data file. Renamed columns should be checked to confirm correct naming if duplicates are present. -
verboseTRUEby default; this and mostmnirsfunctions will return warnings and informational messages which are useful for troubleshooting and data validation. This option can be used to silence those messages.mnirsmessages can be silenced globally for a session by settingoptions(mnirs.verbose = FALSE).
## {mnirs} includes sample files from a few NIRS devices
example_mnirs()
#> [1] "artinis_intervals.xlsx" "moxy_intervals.csv"
#> [3] "moxy_ramp.xlsx" "portamon-oxcap.xlsx"
#> [5] "train.red_intervals.csv"
## partial matching will error if matches multiple
try(example_mnirs("moxy"))
#> Error in example_mnirs("moxy") : ✖ Multiple files match "moxy":
#> ℹ Matching files: "moxy_intervals.csv" and "moxy_ramp.xlsx"
data_raw <- read_mnirs(
file_path = example_mnirs("moxy_ramp"), ## call an example data file
nirs_channels = c(
smo2_left = "SmO2 Live", ## identify and rename channels
smo2_right = "SmO2 Live(2)"
),
time_channel = c(time = "hh:mm:ss"), ## date-time format will be converted to numeric
event_channel = NULL, ## leave blank if unused
sample_rate = NULL, ## if blank, will be estimated from time_channel
add_timestamp = FALSE, ## omit a date-time timestamp column
zero_time = TRUE, ## recalculate time values from zero
keep_all = FALSE, ## return only the specified data channels
verbose = TRUE ## show warnings & messages
)
#> ! Estimated `sample_rate` = 2 Hz.
#> ℹ Define `sample_rate` explicitly to override.
#> Warning: ! Duplicate or irregular `time_channel` samples detected.
#> ℹ Investigate at `time` = 211.59 and 1183.6.
#> ℹ Re-sample with `mnirs::resample_mnirs()`.
## Note the above info message that sample_rate was estimated correctly at 2 Hz ☝
## ignore the warnings about irregular sampling for now, we will resample later
data_raw
#> # A tibble: 2,202 × 3
#> time smo2_left smo2_right
#> <dbl> <dbl> <dbl>
#> 1 0 54 68
#> 2 0.560 54 68
#> 3 1.11 54 66
#> 4 1.66 54 66
#> 5 2.21 54 66
#> 6 2.76 54 66
#> 7 3.31 57 67
#> 8 3.86 57 67
#> 9 4.41 57 67
#> 10 4.96 57 67
#> # ℹ 2,192 more rows📊 Plot mnirs data
mnirs data can be easily viewed by calling plot() (explicitly documented as plot.mnirs()). This generic plot function uses ggplot2 and will work on data frames generated or read by mnirs functions where the metadata contains class = *"mnirs"*.
plot.mnirs
-
dataThis function takes in a data frame of class
mnirsand returns a formattedggplot2plot. -
pointsFALSEby default; a quick way to plot individual points for more precise sample visualisation. -
time_labelsFALSEby default; time values on the x-axis will be plotted as numeric by default.time_labels = TRUEwill instead plot time values ash:mm:ssformat. -
n.breaksDefines the number of breaks plotted on the x-axis, passed to the
ggplot2settings. -
na.omitFALSEby default; missing data (NAs) will be plotted as gaps in the time-series data. Ifna.omit = TRUE,NAs will be omitted from the plotted data, effectively plotting across these gaps, making them less visible, but making the general data trend easier to see if there are lots of missing values.
## note the `time_labels` plot argument to display time values as `h:mm:ss`
plot(
data_raw,
points = FALSE,
time_labels = TRUE,
n.breaks = 5,
na.omit = FALSE
)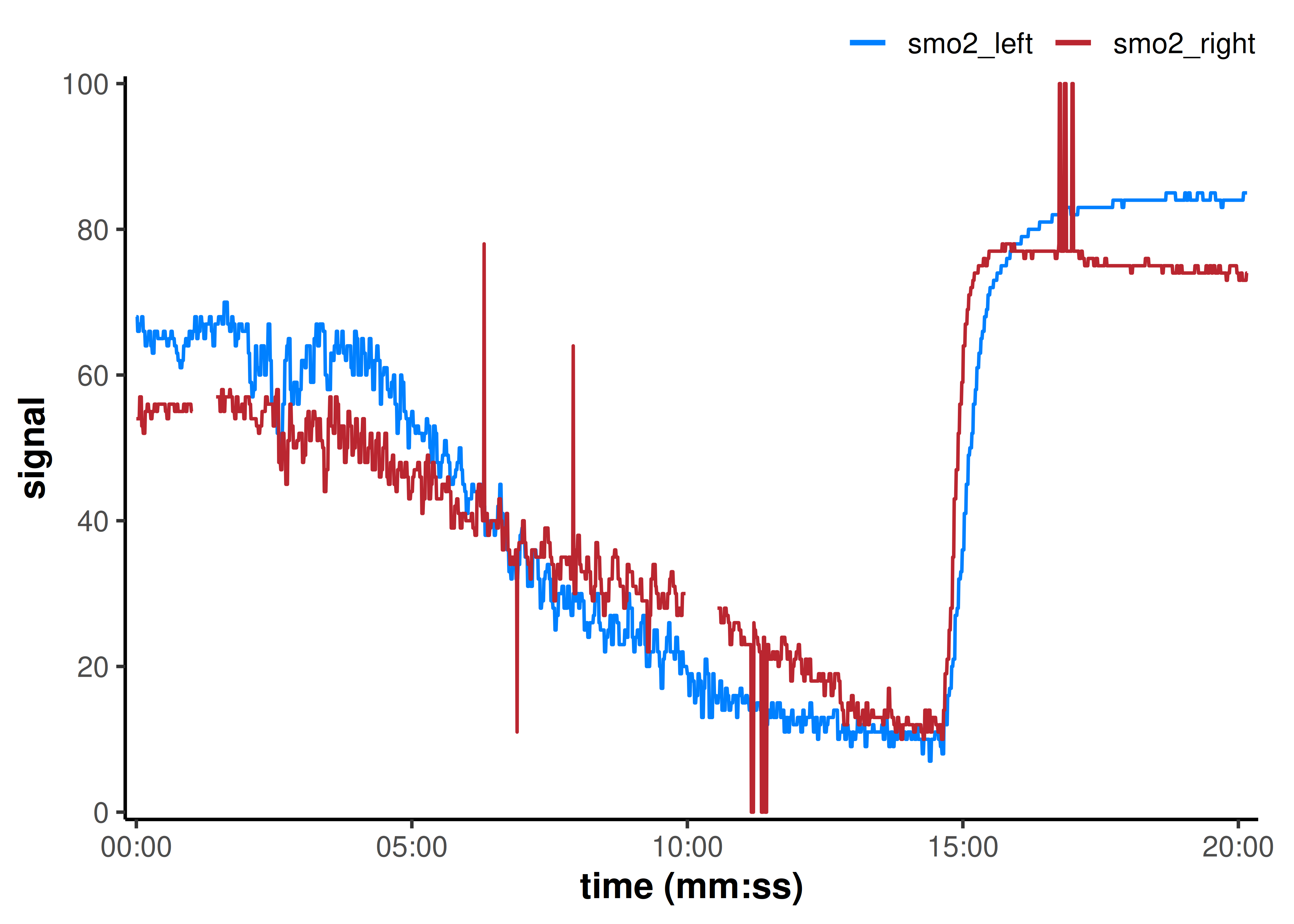
mnirs includes a custom ggplot2 theme and colour palette available with theme_mnirs() and palette_mnirs(). See those documentation references for more details.
🔍 Metadata stored in mnirs data frames
Data frames generated or read by mnirs functions will return class = *"mnirs"* and contain metadata, which can be retrieved with attributes().
Instead of re-defining values like our channel names or sample rate, certain mnirs functions can automatically retrieve them from metadata. They can always be overwritten manually in subsequent functions, or by using a helper function create_mnirs_data().
See create_mnirs_data() for more details about metadata.
⏱️ Resample data
mNIRS devices may have sample rates anywhere from 0.5 to 100+ Hz. Higher sample rates gives greater precision. However, issues can arise with irregular sampling where time samples occur at inconsistent frequencies, are repeated, or are lost. Resampling to a consistent sample rate can be used to regularise timestamps to the nearest time value according to the known data sample rate. This can make subsequent processing steps easier, as well as synchronisation with other devices.
Downsampling can be useful to reduce data size and processing requirement, although should not be used primarily as a data filtering/smoothing strategy (see Digital filtering section below). Upsampling can be done with interpolation over newly created samples, although caution should always be taken with creating new samples without any additional information. New samples created with resampling will be left blank (NA) by default.
resample_mnirs()
-
dataThis function takes in a data frame, applies processing to all channels specified explicitly or implicitly from
mnirsmetadata, then returns the processed data frame.mnirsmetadata will be passed to and from this function.mnirsfunctions are also pipe-friendly for Base R 4.1+ (|>) or magrittr (%>%) pipes to chain operations together (see below). -
time_channelThe column name of time series values in
data. If the data containmnirsmetadata, this channel will be detected automatically, or it can be specified explicitly.
This function processes all columns indata, sonirs_channelsdoes not need to be specified. -
sample_rateThe sample rate (in Hz) of
data. If the data containmnirsmetadata, this channel will be detected automatically, or it can be specified explicitly. -
resample_rateResampling is specified as the desired number of samples per second (Hz). The default
resample_ratewill resample back to the existingsample_rateof the data. This can be useful to accommodate irregular sampling with unequal time values. Linear interpolation is used to resampletime_channelto round values of thesample_rate. -
methodAny new samples created by resampling (i.e. samples fill in between gaps, or up-sampled indices) can be filled in or left blank as
NA. The defaultmethodis to leave new samples blank, and interpolation must be explicitly specified as either “locf” to repeat the last observation carried forward, or “linear” interpolation.
data_resampled <- resample_mnirs(
data_raw, ## blank channels will be retrieved from metadata
time_channel = time, ## channels can be left blank or specified explicitly
sample_rate = NULL, ## blank by default will be retrieved from metadata
resample_rate = 2, ## blank by default will resample to `sample_rate`
method = "linear" ## linear interpolation across resampled indices
)
#> ℹ Output is resampled at 2 Hz.
## note the altered "time" values from the original data frame 👇
data_resampled
#> # A tibble: 2,419 × 3
#> time smo2_left smo2_right
#> <dbl> <dbl> <dbl>
#> 1 0 54 68
#> 2 0.5 54 68
#> 3 1 54 66.4
#> 4 1.5 54 66
#> 5 2 54 66
#> 6 2.5 54 66
#> 7 3 55.3 66.4
#> 8 3.5 57 67
#> 9 4 57 67
#> 10 4.5 57 67
#> # ℹ 2,409 more rowsUse
resample_mnirs()to smooth over irregular or skipped samplesIf we see a warning from
read_mnirs()about duplicated or irregular samples like we saw above, we can useresample_mnirs()to restoretime_channelto a regular sample rate and interpolate across skipped samples.Note: if we perform both of these steps in a piped function call, we will still see the warning appear about irregular sampling at the end of the pipe. This warning is returned by
read_mnirs(). But viewing the data frame can confirm thatresample_mnirs()has resolved the issue.
🧹 Replace local outliers, invalid values, and missing values
We can see some data issues in the plot above, so let’s clean those with a single wrangling function replace_mnirs(), to prepare our data for digital filtering and smoothing.
mnirs tries to include basic functions which work on vector data, and convenience wrappers which combine functionality and can be used on multiple channels in a data frame at once.
Let’s explain the functionality of this data-wide function, and for more details about the vector-specific functions see replace_mnirs().
replace_mnirs()
-
dataThis function takes in a data frame, resamples all data frame columns to the new sample rate, then returns the processed data frame.
mnirsmetadata will be passed to and from this function. -
nirs_channelsSpecify which column names in
datato process, i.e. the response variables. If the data containmnirsmetadata, these channels will be detected automatically. Channels not explicitly specified will be passed through unprocessed to the returned data frame. -
time_channel&sample_rateIf the data contain
mnirsmetadata, these will be detected automatically, or they can be specified explicitly. -
invalid_values,invalid_above, orinvalid_belowSpecific invalid values can be replaced, e.g. if a NIRS device exports
0,100, or some other fixed value when signal recording is in error. If spikes or drops are present in the data, these can be replaced by specifying values above or below which to consider invalid. If left asNULL, no values will be replaced. -
outlier_cutoffLocal outliers can be detected using a cutoff calculated from the local median value. A default value of
3is recommended and corresponds to Pearson’s rule (i.e., ± 3 SD about the local median). If left asNULL, no outliers will be replaced. -
widthorspanLocal outlier detection and median interpolation use a rolling local window specified by one of either
widthorspan.widthdefines a number of samples centred around the local index being evaluated (idx), whereasspandefines a range of time in units oftime_channel. -
methodMissing data (
NA), invalid values, and local outliers specified above can be replaced via interpolation or fill methods; either"linear"interpolation (the default), fill with local"median", or"locf"(“last observation carried forward”).NAs can be passed through to the returned data frame withmethod = "none". However, subsequent processing & analysis steps may return errors whenNAs are present. Therefore, it is good practice to deal with missing data early during data processing. -
verboseAs above, an option to toggle warnings and info messages.
data_cleaned <- replace_mnirs(
data_resampled, ## blank channels will be retrieved from metadata
invalid_values = 0, ## known invalid values in the data
invalid_above = 90, ## remove data spikes above 90
outlier_cutoff = 3, ## recommended default value
width = 7, ## window to detect and replace outliers/missing values
method = "linear" ## linear interpolation over `NA`s
)
plot(data_cleaned, time_labels = TRUE)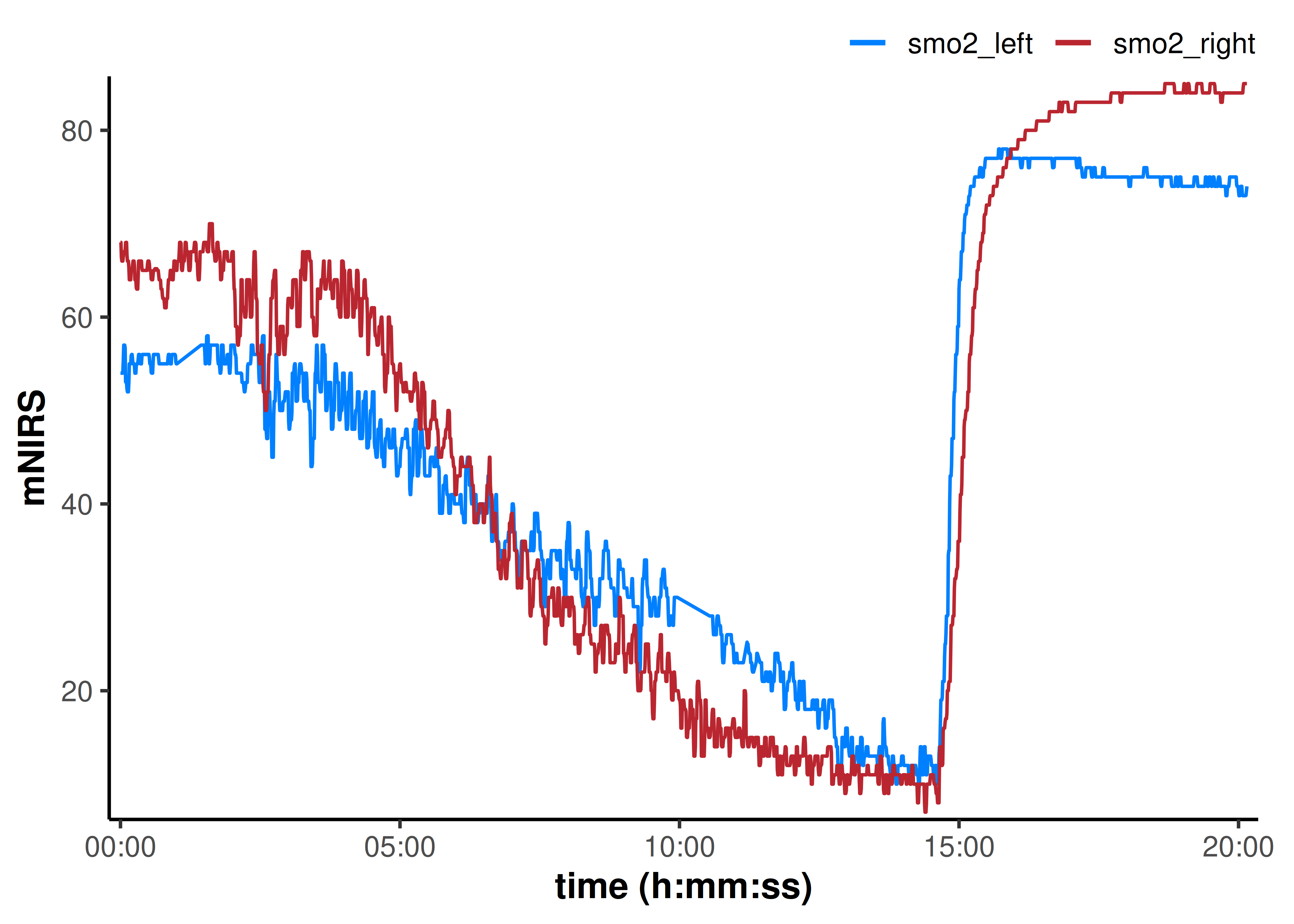
That cleaned up all the obvious data issues.
📈 Digital filtering
To improve the signal-to-noise ratio in our dataset without losing information, we should apply digital filtering to smooth the data.
Choosing a digital filter
There are a few digital filtering methods available in mnirs. Which option is best for you will depend in large part on the sample rate of your data and the frequency of the response signal or phenomenon you are interested in observing.
Choosing filter parameters is an important processing step to improve signal-to-noise ratio and enhance our subsequent interpretations. Over-filtering the data can introduce data artefacts which can negatively influence signal analysis and interpretations, just as much as trying to analyse overly-noisy raw data.
It is perfectly valid to choose a digital filter by iteratively testing filter parameters until the signal or response of interest appears to be visually optimised with minimal data artefacts, to your satisfaction.
We will discuss the process of choosing a digital filter in more depth in another article coming soon.
filter_mnirs()
-
dataThis function takes in a data frame, applies processing to all channels specified, then returns the processed data frame.
mnirsmetadata will be passed to and from this function. -
nirs_channels,time_channel, &sample_rateIf the data contain
mnirsmetadata, these will be detected automatically, or they can be specified explicitly. -
na.rmFALSEby default;filter_mnirs()will return an error if any missing data (NA) are detected in the response variables (nirs_channels). Settingna.rm = TRUEwill ignore theseNAs and pass them through to the returned data frame, but this must be opted into explicitly in this function and elsewhere.
Smoothing-spline
-
method = "smooth_spline"The default non-parametric cubic smoothing spline is often a good first filtering option when exploring the data, and works well over longer time spans. For more rapid and repeated responses, a smoothing-spline may not work as well.
-
sparThe smoothing parameter of the cubic spline will be determined automatically by default, or can be specified explicitly. See
stats::smooth.spline().
Butterworth digital filter
-
method = "butterworth"A Butterworth low-pass digital filter (specified by
type = "low") is probably the most common method used in mNIRS research (whether appropriately or not). For certain applications such as identifying a signal with a known frequency (e.g. cycling/running cadence or heart rate), a pass-band or a different filter type may be better suited. -
orderThe filter order number, specifying the number of passes the filter performs over the data. The default
order = 2will often noticeably improve the filter over a single pass, however, higher orders above ~4can begin to introduce artefacts, particularly at sharp transition points. -
WorfcThe cutoff frequency can be specified either as
W; a fraction (between[0, 1]) of the Nyquist frequency, which is equal to half of thesample_rateof the data. Or asfc; the cutoff frequency in Hz, where this absolute frequency should still be between 0 Hz and the Nyquist frequency. -
typeThe filter type is specified as either
c("low", "high", "stop", "pass").
For filtering vector data and more details about Butterworth filter parameters, see filter_butter().
Moving average
-
method = "moving_average"The simplest smoothing method is a moving average filter applied over a specified number of samples or time span. Commonly, this might be a 5- or 15-second centred moving average.
-
widthorspanMoving average filtering occurs within a rolling local window specified by one of either
widthorspan.widthdefines a number of samples centred around the local index being evaluated (idx), whereasspandefines a range of time in units oftime_channel.
For filtering vector data and more details about the moving average filter, see filter_ma().
Apply the filter
Let’s try a Butterworth low-pass filter, and we’ll specify some empirically chosen filter parameters for these data. See filter_mnirs() for further details on each of these filtering methods and their respective parameters.
data_filtered <- filter_mnirs(
data_cleaned, ## blank channels will be retrieved from metadata
method = "butterworth", ## Butterworth digital filter is a common choice
order = 2, ## filter order number
W = 0.02, ## filter fractional critical frequency `[0, 1]`
type = "low", ## specify a "low-pass" filter
na.rm = TRUE ## explicitly ignore NAs
)
## we will add the non-filtered data back to the plot to compare
plot(data_filtered, time_labels = TRUE) +
geom_line(
data = data_cleaned,
aes(y = smo2_left, colour = "smo2_left"), alpha = 0.4
) +
geom_line(
data = data_cleaned,
aes(y = smo2_right, colour = "smo2_right"), alpha = 0.4
)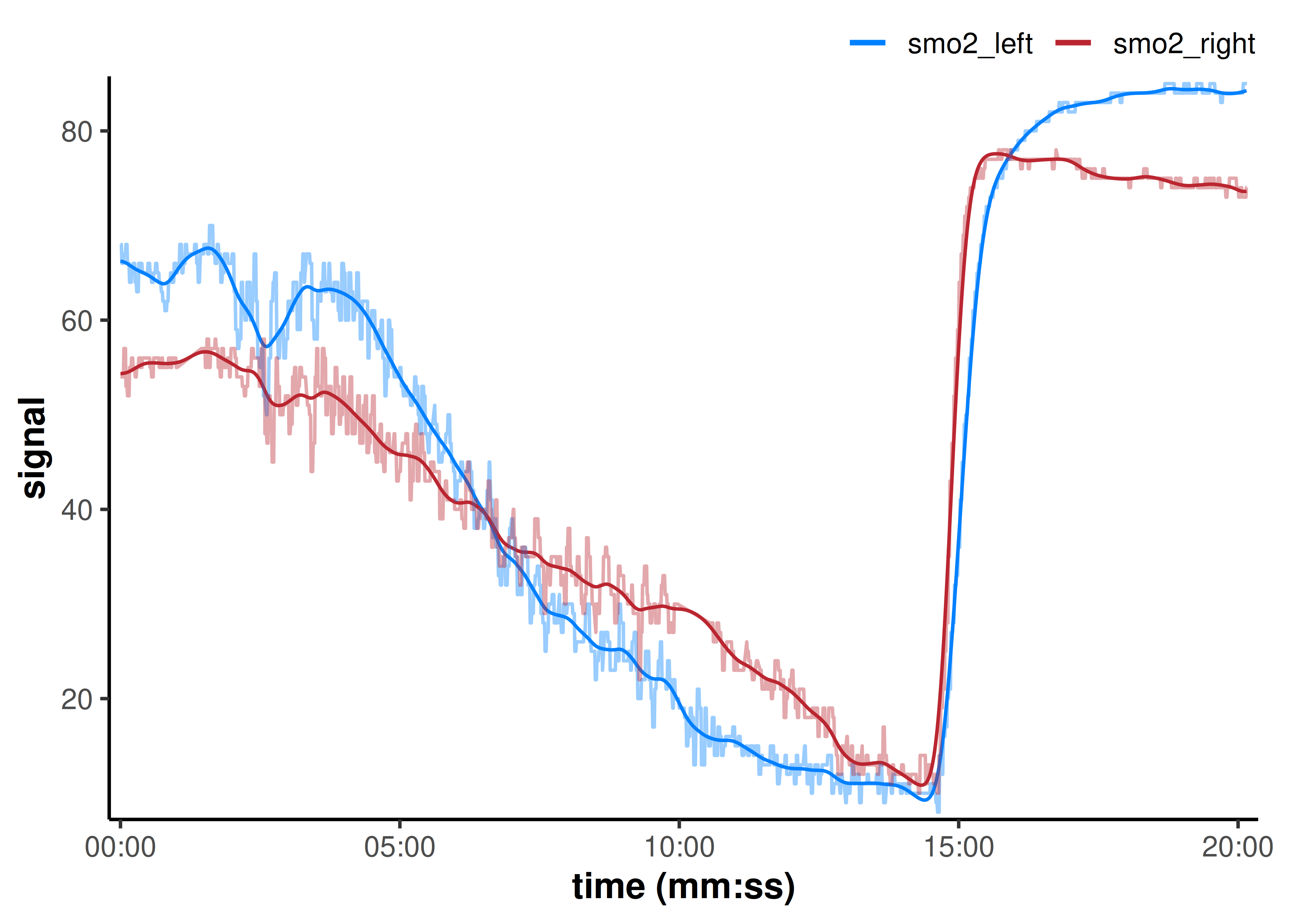
That did a nice job reducing the high-frequency noise in our data, while preserving the sharp edges, such as the rapid reoxygenation after maximal exercise.
⚖️ Shift and rescale data
NIRS values are not measured on an absolute scale (arguably not even percent (%) saturation/SmO2). Therefore, we may need to adjust or calibrate our data to normalise NIRS signal values between muscle sites, individuals, trials, etc. depending on our intended comparison.
For example, we may want to set our mean baseline value to zero for all NIRS signals at the start of a recording. Or we may want to compare signal kinetics (the rate of change or time course of a response) after rescaling signal amplitudes to a common dynamic range.
These functions allow us to either shift NIRS values up or down while preserving the dynamic range (the absolute amplitude from minimum to maximum values) of our NIRS channels, or rescale the data to a new dynamic range with larger or smaller amplitude.
We can group NIRS channels together to preserve the absolute and relative scaling among certain channels, and modify that scaling between other groups of channels.
shift_mnirs()
-
dataThis function takes in a data frame, applies processing to all channels specified, then returns the processed data frame.
mnirsmetadata will be passed to and from this function. -
nirs_channelsUnlike previous functions,
nirs_channelsshould provided as a list (e.g.list(c(A, B), c(C))). Each list item represents a group to be shifted together to a common scale. Separate list items will be shifted to separate scales. The relative scaling between channels will be preserved within each group, but lost between groups.nirs_channelsshould be specified explicitly to ensure the intended grouping structure is returned. The defaultmnirsmetadata will group all NIRS channels together. -
time_channelIf the data contain
mnirsmetadata, this channel will be detected automatically, or it can be specified explicitly. -
toorbyThe shift amplitude can be specified by either shifting signals
toa new value, or shifting signalsbya fixed amplitude, given in units of the NIRS signals. -
widthorspanShifting can be performed on the mean value within a window specified by one of either
widthorspan.widthdefines a number of samples centred around the local index being evaluated (idx), whereasspandefines a range of time in units oftime_channel.
e.g.width = 1will shift from the single minimum sample, whereasspan = 1would shift from the minimum mean value of all samples within one second. -
positionSpecifies the reference point used to shift the data; either
"min","max", or"first"sample(s).
For this data set, we want to shift each NIRS channel so that the mean of the 2-minute baseline is equal to zero, which would then give us a change in deoxygenation from baseline during the incremental cycling protocol.
tidyverse-style channel name specification
This is a good time to note that here and in most
mnirsfunctions, data channels can be specified using tidyverse-style naming. Data frame column names can be specified either with quotes as a character string ("smo2"), or as a direct symbol (smo2).tidyselect support functions such as
starts_with(),matches()can also be used.
data_shifted <- shift_mnirs(
data_filtered, ## un-grouped nirs channels to shift separately
nirs_channels = list(smo2_left, smo2_right), ## 👈 channels grouped separately
to = 0, ## NIRS values will be shifted to zero
span = 120, ## shift the *first* 120 sec of data to zero
position = "first"
)
plot(data_shifted, time_labels = TRUE) +
geom_hline(yintercept = 0, linetype = "dotted")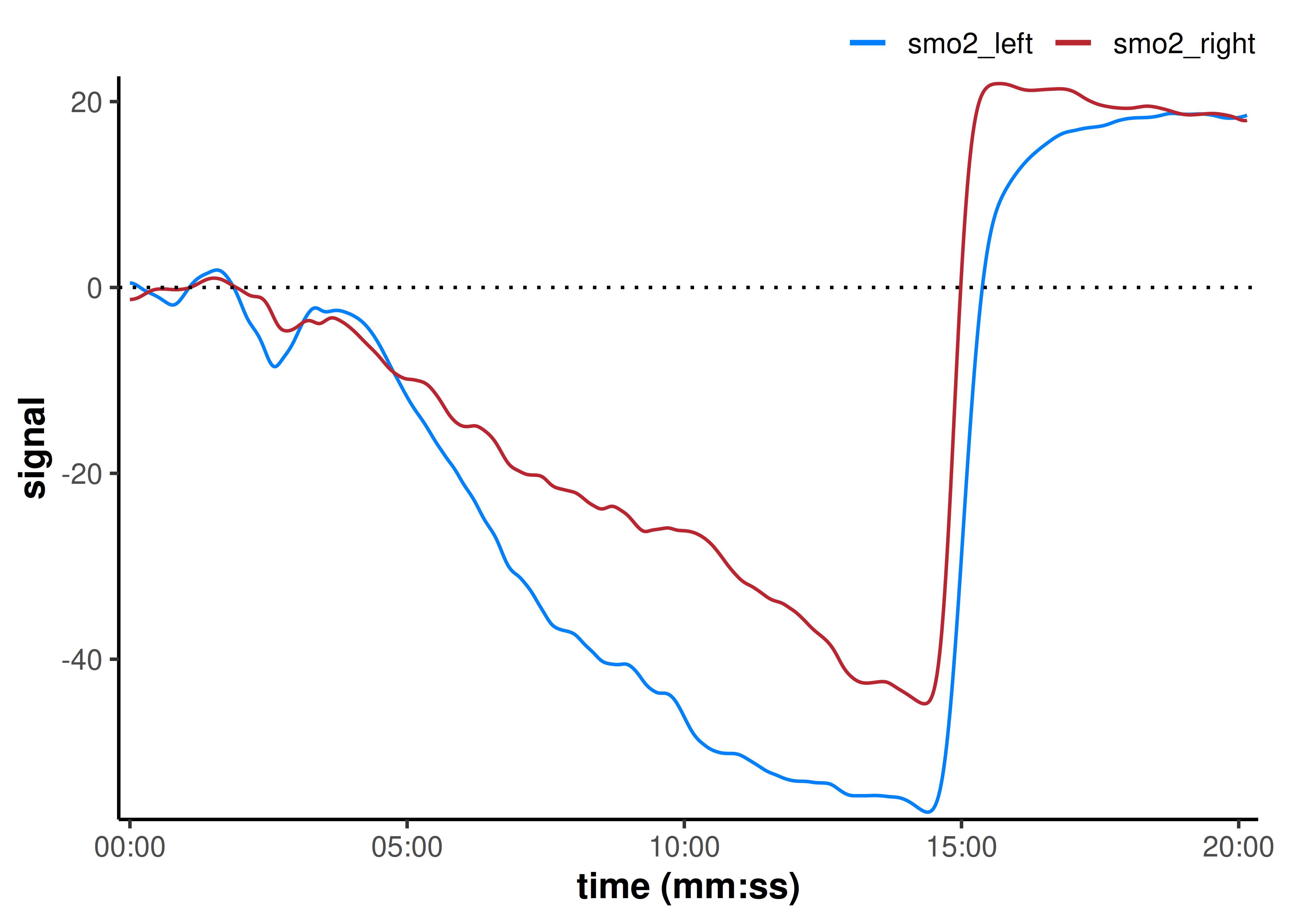
Before shifting, the minimum (end of exercise) values for smo2_left and smo2_right were similar, but the starting baseline values were different.
When we shift both baseline values to zero, however, we can see that the smo2_left signal has a smaller deoxygenation amplitude compared to the smo2_right signal, and a (slightly) greater hyperaemic reoxygenation peak.
We have to consider how our assumptions and processing decisions will influence our interpretations; by shifting both starting values, we are normalising for, and implicitly assuming that the baseline represents the same starting condition for the tissues in both legs.
This may or may not be an appropriate assumption for your research question; for example, this may be appropriate when we are more interested in the relative change (delta) in each leg during an intervention or exposure (often referred to as "∇SmO2"), but not if we were interested in asymmetries that could influence SmO2 at rest.
rescale_mnirs()
We may also want to rescale our data to a new dynamic range, changing the signal amplitude.
-
dataThis function takes in a data frame, applies processing to all channels specified, then returns the processed data frame.
mnirsmetadata will be passed to and from this function. -
nirs_channelsUnlike previous functions,
nirs_channelsshould provided as a list (e.g.list(c(A, B), c(C))). Each list item represents a group to be rescale together to a common range. Separate list items will be rescaled to separate ranges. The relative scaling between channels will be preserved within each group, but lost between groups.nirs_channelsshould be specified explicitly to ensure the intended grouping structure is returned. The defaultmnirsmetadata will group all NIRS channels together. -
rangeSpecifies the new dynamic range in the form
c(min, max). For example, if we want to calibrate each NIRS signal to its own observed ‘functional range’ during exercise, we could rescale them to 0-100%.
data_rescaled <- rescale_mnirs(
data_filtered, ## un-grouped nirs channels to rescale separately
nirs_channels = list(smo2_left, smo2_right), ## 👈 channels grouped separately
range = c(0, 100) ## rescale to a 0-100% functional exercise range
)
plot(data_rescaled, time_labels = TRUE) +
geom_hline(yintercept = c(0, 100), linetype = "dotted")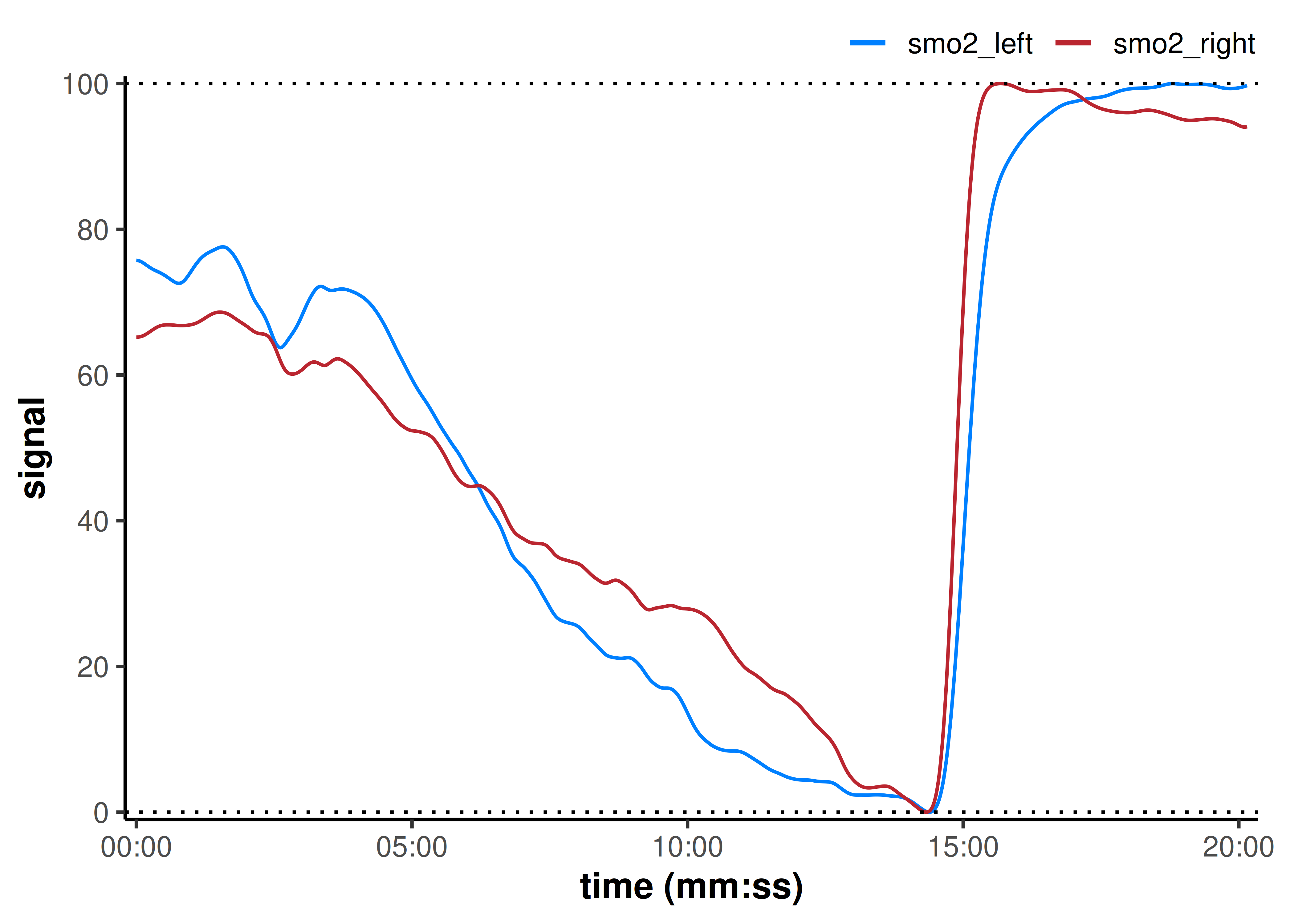
Here, our assumption is that during a maximal exercise task, the minimum and maximum values represent the functional capacity of each tissue volume being observed. So we rescale the functional dynamic range in each leg.
By normalising this way, we might lose meaningful differences captured by the different amplitudes between smo2_left and smo2_right, but we might be more interested in the trend or time course of each response.
Our interpretation may be that smo2_right appears to start at a slightly higher percent of its functional range, deoxygenates faster toward a minimum, and reaches a quasi-plateau near maximal exercise. While smo2_left deoxygenates slightly slower and continues to deoxygenate until maximal task tolerance.
Additionally, the left leg reoxygenates slightly faster than the right during recovery. This might, for example, indicate exercise capacity differences between the limbs (although these differences are marginal and only discussed as representative for influence on interpretations).
🔀 Pipe-friendly functions
Most mnirs functions can be piped together using Base R 4.1+ (|>) or magrittr (%>%). The entire pre-processing stage can easily be performed in a sequential pipe.
To demonstrate this, we’ll read a different example file recorded with Train.Red FYER muscle oxygen sensor and pipe it through each processing stage straight to a plot.
This is also a good time to demonstrate how to use the global mnirs.verbose option to silence all warning & information messages, for example when dealing with a familiar dataset. We recommend leaving verbose = TRUE by default whenever reading and exploring a new file.
options(mnirs.verbose = FALSE)
nirs_data <- read_mnirs(
example_mnirs("train.red"),
nirs_channels = c(
smo2_left = "SmO2 unfiltered",
smo2_right = "SmO2 unfiltered"
),
time_channel = c(time = "Timestamp (seconds passed)"),
zero_time = TRUE
) |>
resample_mnirs(method = "linear") |> ## default settings will resample to the same `sample_rate`
replace_mnirs(
invalid_above = 73,
outlier_cutoff = 3,
span = 7
) |>
filter_mnirs(
method = "butterworth",
order = 2,
W = 0.02,
na.rm = TRUE
) |>
shift_mnirs(
nirs_channels = list(smo2_left, smo2_right), ## 👈 channels grouped separately
to = 0,
span = 60,
position = "first"
) |>
rescale_mnirs(
nirs_channels = list(c(smo2_left, smo2_right)), ## 👈 channels grouped together
range = c(0, 100)
)
plot(nirs_data, time_labels = TRUE)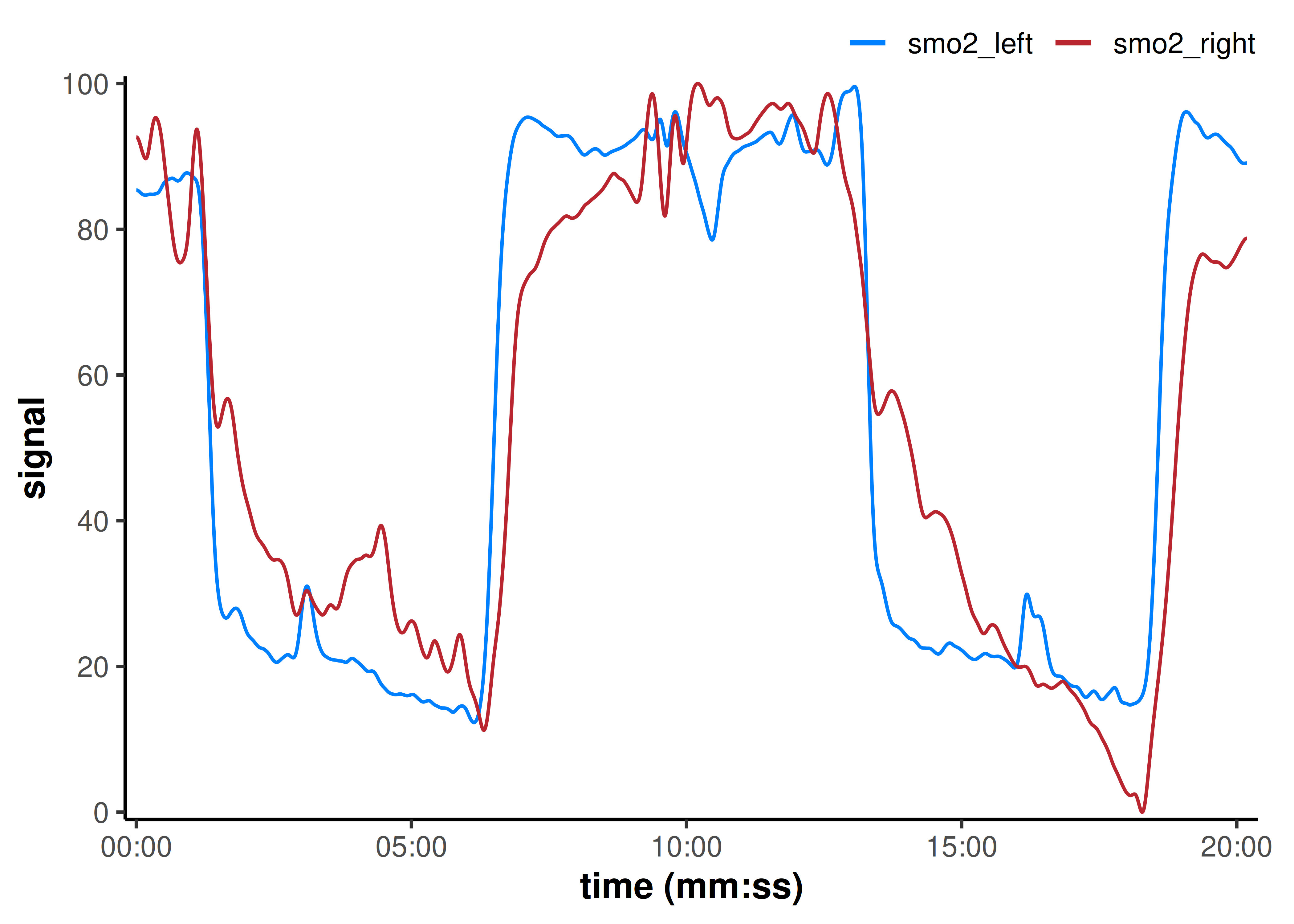
We have two exercise intervals in this data set. Let’s demonstrate some of the common analysis methods currently available with mnirs.
🧮 Interval extraction
After the NIRS signal has been cleaned and filtered, it should be ready for further processing and analysis.
mnirs is under development to include functionality for processing discrete intervals and events, e.g. reoxygenation kinetics, slope calculations for post-occlusion microvascular responsiveness, and critical oxygenation breakpoints.
extract_intervals()
Often we will need to locate and extract smaller intervals from a NIRS recording for further processing and analysis. For example, if we want to extract the last three minutes from repeated exercise trials.
We may have marked event labels or have incremental lap numbers in the specified event_channel, or we may simply have to manually specify the event markers by time_channel value or row number in the data frame. See extract_intervals() for more details.
-
dataThis function takes in a data frame, applies processing to all channels specified, then returns a list of processed data frames.
mnirsmetadata will be passed to and from this function. -
nirs_channelsIf returning a list of “distinct” intervals (see
event_groupsbelow),nirs_channelsdoes not have to be specified, as no channels are processed.
Only when “ensemble”-averaging,nirs_channelsshould be specified by providing a list of column names (e.g.list(c(A, B), c(A))), where each list item specifies the channels to be ensemble-averaged within the respective group (ensemble-groups are specified byevent_groupsbelow), in the order in which they are returned. The defaultmnirsmetadata will ensemble-average allnirs_channels. -
time_channel,event_channel, &sample_rateIf the data contain
mnirsmetadata, these will be detected automatically, or they can be specified explicitly. -
start&endInterval boundaries are specified using helper functions:
by_time()fortime_channelvalues;by_sample()for sample indices (row numbers);by_label()forevent_channellabels; orby_lap()forevent_channellap integers. Provide bothstartandendto define precise intervals, or providestartalone withspanto extract windows around events. Multiple values can be passed to extract multiple intervals. -
event_groupsEvents can be extracted and returned as a list of
"distinct"intervals, or"ensemble"-averaged into a single data frame. Custom grouping structure for ensemble-averaging can be specified by event number, in order of appearance within the original data.
e.g.event_groups = list(c(1, 2), c(3, 4))would return a list of two intervals, each ensemble-averaged from the respective events, in sequential order from the original data. -
spanWhen only
start(orend) is provided,spanspecifies a time window in units oftime_channelasc(before, after), where positive values indicate time after the event and negative values indicate time before. When bothstartandendare provided,spanshifts boundaries additively:span[1]adjusts starts,span[2]adjusts ends.
A list of uniquespanvectors can be specified for each interval, otherwise a singlespanvector will be recycled to all intervals. -
zero_timeFALSEby default; becausetime_channelvalues within each returned interval likely start at a non-zero value,zero_time = TRUEwill re-calculate time starting from zero at the specified event, so the start time (or however an event is specified) will become0.
When ensemble-averaging across intervals, time will always be re-calculated from0, since the time values have lost their meaning.
## return each interval independently with `event_groups = "distinct"`
distinct <- extract_intervals(
nirs_data, ## channels blank for "distinct" grouping
start = by_time(348, 1064), ## manually identified interval start times
end = by_time(458, 1174), ## interval end time (start + 150 sec)
event_groups = "distinct", ## return a list of data frames for each (2) event
span = c(0, 0), ## no boundary modification
zero_time = FALSE ## return original time values
)
plot(distinct, time_labels = TRUE)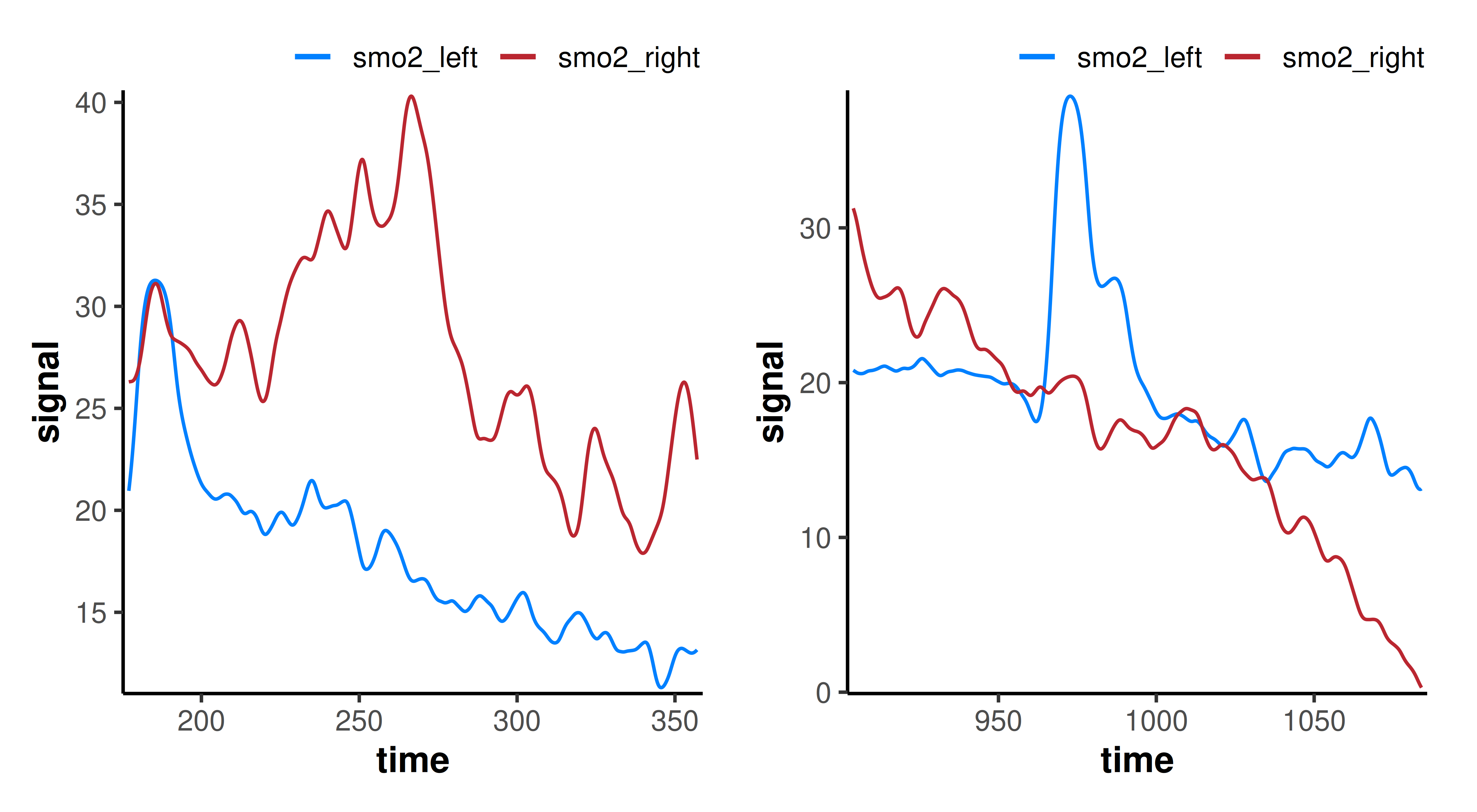
## ensemble average both intervals with `event_groups = "ensemble"`
ensemble <- extract_intervals(
nirs_data, ## channels recycled to all intervals by default
nirs_channels = c(smo2_left, smo2_right),
start = by_time(368, 1084), ## alternatively specify start times + span
event_groups = "ensemble", ## ensemble-average across two intervals
span = c(-20, 90), ## span recycled to all intervals by default
zero_time = TRUE ## re-calculate common time to start from `0`
)
plot(ensemble, time_labels = TRUE) +
geom_vline(xintercept = 0, linetype = "dotted")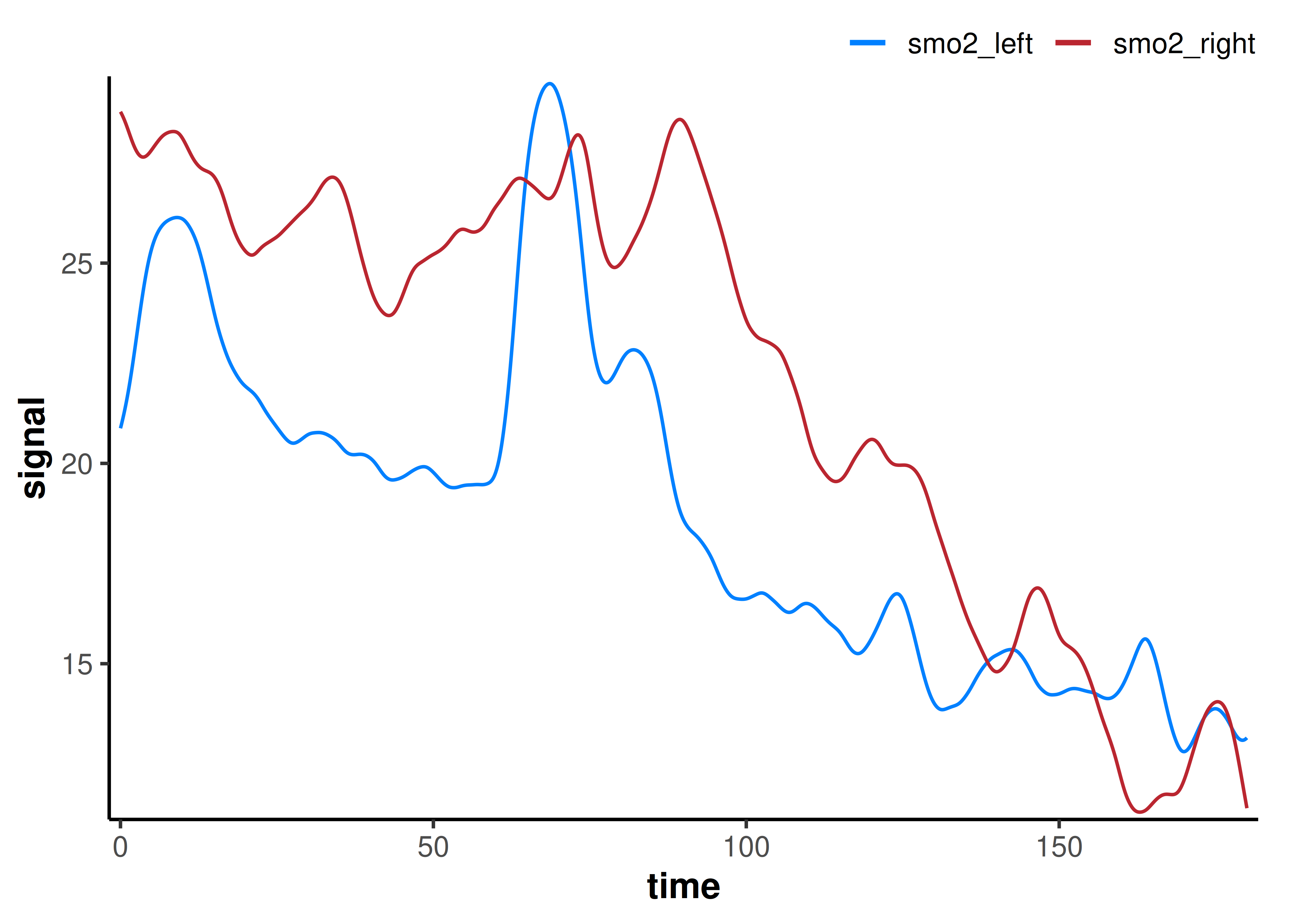
The ensemble-averaged kinetics capture a more representative ‘average’ response during the two interval deoxygenation events.
Ensemble-averaging can help mitigate data dropouts and other quality issues from measurement error or biological variability. For example, ensemble-averaging is common for evaluating systemic VO2 kinetics, where responses can be highly variable trial to trial.
Conclusion
This vignette walks through the core functionality of mnirs to read, clean, and pre-process data in preparation for analysis.
Future development and articles will cover standardised analysis methods, including deoxygenation & reoxygenation kinetics and critical oxygenation breakpoint analysis.