Apply digital filtering/smoothing to numeric vector data within a data frame using either:
A cubic smoothing spline.
A Butterworth digital filter.
A simple moving average.
Usage
filter_mnirs(
data,
nirs_channels = NULL,
time_channel = NULL,
method = c("smooth_spline", "butterworth", "moving_average"),
na.rm = FALSE,
verbose = TRUE,
...
)Arguments
- data
A data frame of class "mnirs" containing time series data and metadata.
- nirs_channels
A character vector giving the names of mNIRS columns to operate on. Must match column names in
dataexactly.If
NULL(default), thenirs_channelsmetadata attribute ofdatais used.
- time_channel
A character string naming the time or sample column. Must match a column name in
dataexactly.If
NULL(default), thetime_channelmetadata attribute ofdatais used.
- method
A character string indicating how to filter the data (see Details).
"smooth_spline"Fits a cubic smoothing spline.
"butterworth"Uses a centred Butterworth digital filter.
"moving_average"Uses a centred moving average filter.
- na.rm
Logical; default is
FALSE, propagates anyNAs to the returned vector. IfTRUE, ignoresNAs and processes available valid samples within the local window. May return errors or warnings. (see Details).- verbose
Logical. Default is
TRUE. Display or silence (ifFALSE) warnings and information messages helpful for troubleshooting. Ad global default can be set viaoptions(mnirs.verbose = FALSE).- ...
Additional method-specific arguments must be specified (see Details).
Value
A tibble of class "mnirs" with metadata
available with attributes().
Details
method = "smooth_spline"
Aliases: method = c("smooth spline", "spline")
Applies a non-parametric cubic smoothing spline from
stats::smooth.spline(). Smoothing is defined by the parameter spar,
which can be left as NULL and automatically determined via penalised
log likelihood. This usually works well for responses occurring on the
order of minutes or longer. spar can be specified typically, but not
necessarily, in the range spar = [0, 1].
Additional arguments (...) accepted when method = "smooth_spline":
sparA numeric smoothing parameter passed to
stats::smooth.spline(). IfNULL(default), automatically determined via penalised log likelihood.
method = "butterworth"
Aliases: method = c("butter")
Applies a centred (two-pass symmetrical) Butterworth digital filter
from signal::butter() and signal::filtfilt().
Filter type defines how the desired signal frequencies are either
passed or rejected from the output signal. Low-pass and high-pass
filters allow only frequencies lower or higher than the cutoff
frequency, respectively to be passed through to the output signal.
Stop-band defines a critical range of frequencies which are rejected
from the output signal. Pass-band defines a critical range of
frequencies which are passed through as the output signal.
The filter order (number of passes) is defined by order, typically
in the range order = [1, 10]. Higher filter order tends to capture
more rapid changes in amplitude, but also causes more distortion
around those change points in the signal. General advice is to use
the lowest filter order which sufficiently captures the desired rapid
responses in the data.
The critical (cutoff) frequency can be defined by W, a numeric value
for low-pass and high-pass filters, or a two-element vector
c(low, high) defining the lower and upper bands for stop-band
and pass-band filters. W represents the desired fractional cutoff
frequency in the range W = [0, 1], where 1 is the Nyquist
frequency, i.e., half the sample_rate of the data in Hz.
Alternatively, the cutoff frequency can be defined by fc and
sample_rate together. fc represents the desired cutoff frequency
directly in Hz, and sample_rate is the sample rate of the recorded data
in Hz. Where W = fc / (sample_rate / 2).
Only one of either W or fc should be defined. If both are
defined, W will be preferred over fc.
Additional arguments (...) accepted when method = "butterworth":
orderAn integer for the filter order (default
2).WA numeric fractional cutoff frequency within
[0, 1]. One of eitherWorfcmust be specified.fcA numeric absolute cutoff frequency in Hz. Used with
sample_rateto computeW.sample_rateA numeric sample rate in Hz. Will be taken from metadata or estimated from
time_channelif not defined.typeA character string specifying filter type, one of:
c("low", "high", "stop", "pass")("low"is the default).edgesA character string specifying the edge padding, one of:
c("rev", "rep1", "none")("rev"is the default). Seefilter_butter().
method = "moving_average"
Aliases: method = c("moving average", "ma")
Applies a centred (symmetrical) moving average filter in a local
window, defined by either width as the number of samples around
idx between [idx - floor(width/2), idx + floor(width/2)]. Or by
span as the timespan in units of time_channel between
[t - span/2, t + span/2].
Additional arguments (...) accepted when method = "moving_average":
widthorspanEither an integer number of samples, or a numeric time duration in units of
time_channelwithin the local window. One of eitherwidthorspanmust be specified.partialLogical;
FALSEby default, only returns values where a full window of valid (non-NA) samples are available. IfTRUE, ignoresNAand allows calculation over partial windows at the edges of the data.
Missing values
Missing values (NA) in nirs_channels will cause an error for
method = "smooth_spline" or "butterworth", unless na.rm = TRUE.
Then NAs will be ignored and passed through to the returned data.
For method = "moving_average", na.rm controls whether NAs within
each local window are either propagated to the returned vector when
na.rm = FALSE (the default), or ignored before processing if
na.rm = TRUE.
Examples
## read example data and clean for outliers
data <- read_mnirs(
file_path = example_mnirs("moxy_ramp"),
nirs_channels = c(smo2 = "SmO2 Live"),
time_channel = c(time = "hh:mm:ss"),
verbose = FALSE
) |>
replace_mnirs(
invalid_values = c(0, 100),
outlier_cutoff = 3,
width = 7,
verbose = FALSE
)
data
#> # A tibble: 2,202 × 2
#> time smo2
#> <dbl> <dbl>
#> 1 0 54
#> 2 0.560 54
#> 3 1.11 54
#> 4 1.66 54
#> 5 2.21 54
#> 6 2.76 54
#> 7 3.31 57
#> 8 3.86 57
#> 9 4.41 57
#> 10 4.96 57
#> # ℹ 2,192 more rows
data_filtered <- filter_mnirs(
data, ## blank channels will be retrieved from metadata
method = "butterworth", ## Butterworth digital filter is a common choice
order = 2, ## filter order number
W = 0.02, ## filter fractional critical frequency `[0, 1]`
type = "low", ## specify a "low-pass" filter
na.rm = TRUE ## explicitly ignore NAs
)
## note the smoothed `smo2` values
data_filtered
#> # A tibble: 2,202 × 2
#> time smo2
#> <dbl> <dbl>
#> 1 0 54.5
#> 2 0.560 54.5
#> 3 1.11 54.5
#> 4 1.66 54.5
#> 5 2.21 54.5
#> 6 2.76 54.5
#> 7 3.31 54.5
#> 8 3.86 54.5
#> 9 4.41 54.5
#> 10 4.96 54.5
#> # ℹ 2,192 more rows
# \donttest{
if (requireNamespace("ggplot2", quietly = TRUE)) {
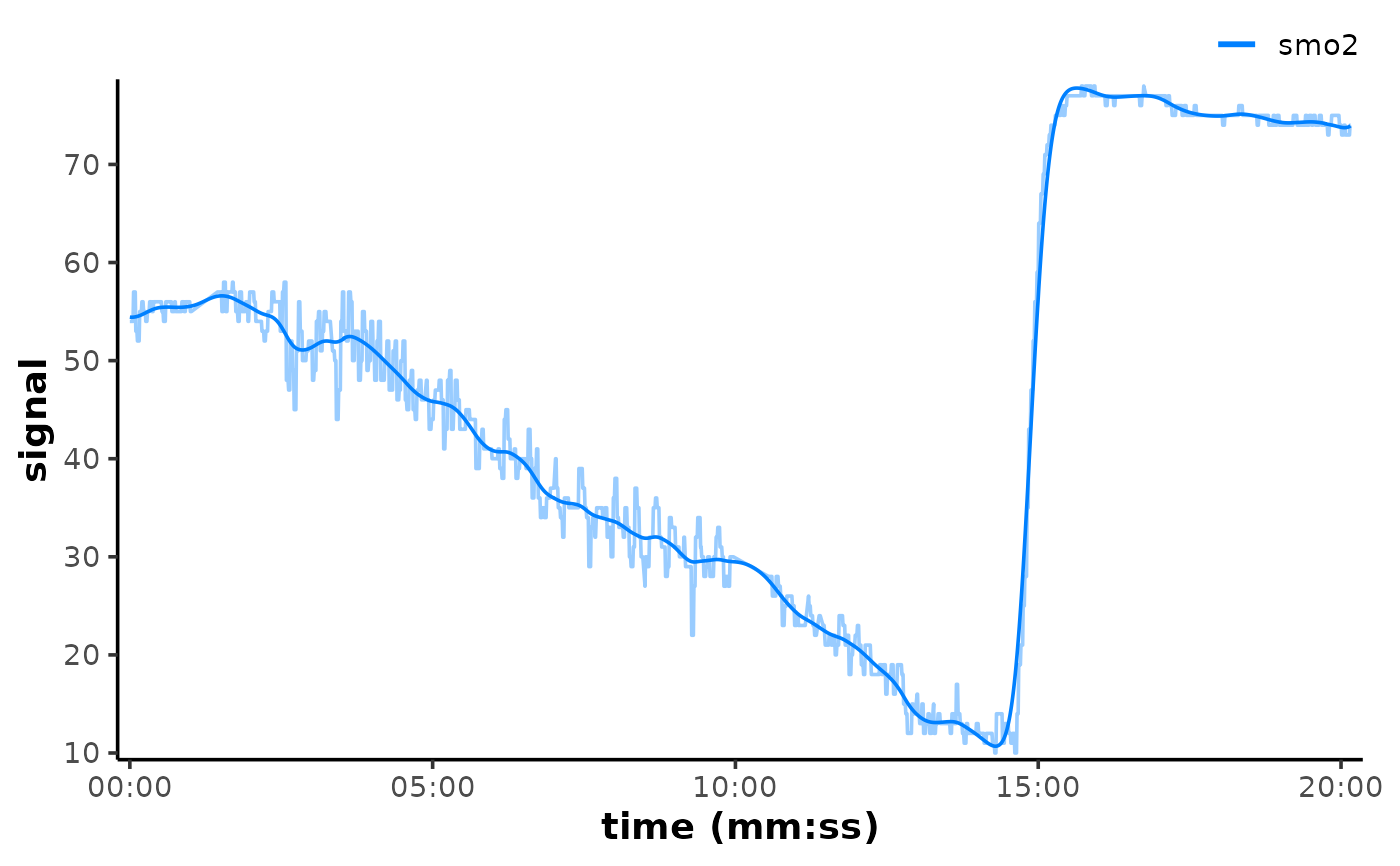
## plot filtered data and add the raw data back to the plot to compare
plot(data_filtered, time_labels = TRUE) +
ggplot2::geom_line(
data = data,
ggplot2::aes(y = smo2, colour = "smo2"), alpha = 0.4
)
}
 # }
# }